 That being said, the need to include graphic figures in order to display sequence logos has perpetuated the use of consensus sequences in scientific manuscripts, even though they fail to convey information on both conservation and frequency. Hence, a sequence logo should be used preferentially whenever possible. As a result, compared to a sequence logo, the consensus logo omits information (the relative contribution of each character to the conservation of that position in the motif/alignment). 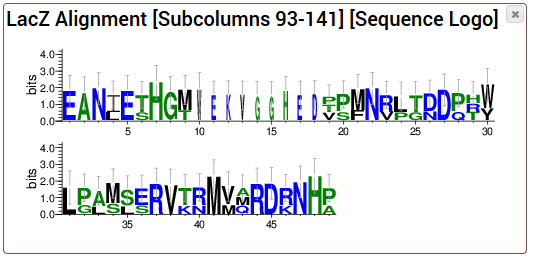
As described above, the consensus logo is a cross between sequence logos and consensus sequences. The main, and obvious, advantage of consensus logos over sequence logos is their ability to be embedded as text in any Rich Text Format supporting editor/viewer and, therefore, in scientific manuscripts. Instead of a stack made of several characters, denoting the relative frequency of each character, the consensus logo depicts the degree of conservation of each position using the height of the consensus character at that position.Ī consensus logo for the LexA-binding motif of several Gram-positive species. However, a consensus logo displays only conservation information, and not explicitly the frequency information of each nucleotide or amino acid at each position. Like a sequence logo, a consensus logo is created from a collection of aligned protein or DNA/RNA sequences and conveys information about the conservation of each position of a sequence motif or sequence alignment The information content (y-axis) of position \displaystyle is the number of sequences in the alignment.Ī consensus logo is a simplified variation of a sequence logo that can be embedded in text format. Sequence logos can be used to represent conserved DNA binding sites, where transcription factors bind. The height of the entire stack of residues is the information measured in bits. Different residues at the same position are scaled according to their frequency. The sequence logo will show how well residues are conserved at each position: the higher the number of residues, the higher the letters will be, because the better the conservation is at that position. A sequence logo can then be created from the conserved multiple sequence alignment. To create sequence logos, related DNA, RNA or protein sequences, or DNA sequences that have common conserved binding sites, are aligned so that the most conserved parts create good alignments. The total height of the letters depicts the information content of the position, in bits. The relative sizes of the letters indicate their frequency in the sequences. 1990 Oct 25 18( 20):6097-100.A sequence logo consists of a stack of letters at each position. Schneider TD, Stephens RM., Sequence logos: a new way to display consensus sequences., Nucleic Acids Res. If you need that sort of data, calculate it from the alignment directly. So don't put too much weight on the bit values. However, you should think of logos as a useful visual aid, and not a way of getting rigorous mathematical information about a sequence. In the one you show, the scale is from 0 to 0.2, but it's the same principle. 
So yes, 2bits is the highest value you will see on the y-axis of a DNA sequence logo (proteins are a different story since there are more possibilities). Is needed to describe a position in a binding site that contains only purines,īut two bits are needed to describe a position that always contains adenine. Sometimes A and sometimes G), only one question suffices since a two out ofįour choice is equivalent to a one out of two choice. (If the answers to both questionsĪre "no", it must be a T.) Furthermore, if a position contains two bases (e.g. Information since two yes-no questions need to be answered: "Is it A or G?" Question needs to be answered: "Is it heads?". For example, to communicate the result ofĪ coin-ip to someone requires 1 bit of information because only one yes-no To choose one symbol or state from two equally likely possibilities The importance of a particular position in a binding site is more clearlyĪnd consistently given by the information required to describe the pattern This is explained in the original paper describing the logos (emphasis mine): 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
